Gemma Church explains how Abberior Instruments’ super-resolution microscopes are tackling Covid-19
Viruses raise many challenges from both an imaging and public health perspective, where we want to observe how they impact the human body to reduce their spread. If the recent Sars-Cov-2 pandemic is anything to go by, we still have a lot of work to do in this area – and the field of super-resolution microscopy is now helping researchers keep pace with the complex and dynamic field of virology.
In fact, microscopy-based experimental approaches have a ‘central role’ to play in the field of modern biomedical research, according to Vibor Laketa, head of the Infectious Disease Imaging Platform at the Center for Integrative Infectious Diseases Research (CIID), who explained: ‘A comprehensive understanding of host-pathogen interactions requires quantitative assessment of molecular events across a wide range of spatiotemporal scales and organisational complexities. Due to recent technical developments, this is currently only achievable with microscopy.’
‘Thus, we believe that modern microscopy technology is currently uniquely positioned to propel infectious disease research to a new frontier,’ Laketa added.
The CIID is a state-of-the art research facility of the Department for Infectious Diseases at the University Hospital Heidelberg, Germany. It works across multiple research fields, including molecular biology, cell biology and immunology of pathogen-host interactions with a particular emphasis on their visualisation at maximal spatial-temporal resolution.
The mission of the CIID is clear, to investigate major human pathogens, define their pathogenic principles, and translate this work into the development of novel therapeutic and preventive strategies.
The Infectious Disease Imaging Platform (IDIP) is one research unit at the CIID. The team employs a reductionist experimental approach, which can capture and quantitatively examine individual molecular events. Such events can reveal the differences between the health and the disease state. This information can then be incorporated into mathematical models together with observations made using population-based recordings, which ultimately leads to a comprehensive understanding of the underlying patho-physiological processes.
The technical challenges to observe and quantify these stochastic molecular events are substantial. So, IDIP is equipped with a comprehensive range of bioimaging technologies that allow researchers to study host-pathogen interactions at all spatio-temporal scales – for everything from microscopic molecules and individual viral particles all the way up to macroscopic-sized whole organs, organoids and whole, small organisms.
The IDIP infrastructure is organised in 14 interconnected microscopy rooms across a 250m2 floor space. Both cryoCLEM microscopy techniques (where a combination of low-temperature fluorescence and electron microscopy is used) and super-resolution technologies are used, as well as other techniques including spinning disc microscopy, selective plane illumination microscopy (SPIM) and multiphoton microscopy.
Laketa explained: ‘As pathogens such as viruses are typically of sizes below the diffraction limit of visible light, technologies such as cryoCLEM and super-resolution allowed major insights in molecular structures of individual microorganisms as well as host-pathogen interactions at the spatial scale below 200nm and tackled questions such as what does the virus look like, how does it change upon cell entry and exit, how does the cellular ultrastructure of the host change upon pathogen encounter?
‘SPIM and multiphoton technologies allow the investigation of dynamic host-pathogen interactions that are happening in complex model systems and provided answers to basic questions such as how do pathogens spread through the organism, which cells in a multicellular organism are infected, why are pathogens making people sick?’ Laketa said.
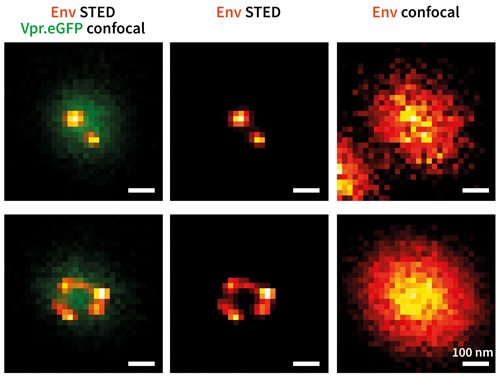
Env protein distribution (red/yellow) on single HIV-1 particles (green). Vpr. eGFP was used as a marker for HIV- 1 particles. Env clusters on the virus particle surface were labelled by Fab fragments coupled to STAR635P. Only with super-resolution microscopy can single Env clusters be revealed (compare middle column (STED) and right column (confocal)). Credit: Jakub Chojnacki
Covid-19 reaction
Like many working in the field of virology, the IDIP research team has been asked what it can do to fight the current Sars-Cov-2 pandemic. In response, the researchers are developing a microscopy-based method to measure levels of Sars-Cov-2 specific antibodies in human serum, which is the clear, yellowish coloured fluid found in blood.
Using this technique, researchers can gain a better understanding of the human body’s immune response against the virus and assess the levels of the pathogen in the population.
The team has already tested more than 5,000 people in the Baden-Württemberg state of Southwest Germany using this technique. ‘Having access to advanced microscopy infrastructure under BSL2 and BSL3 biosafety level and expert personnel was essential for execution of this study. It also shows how scientific infrastructure typically used in academic settings can be rapidly deployed to address emergent medical and public health demands,’ Laketa explained.
The team also characterised pathological changes in cellular organelle structures in the presence of the Sars-Cov-2 infection using stimulated emission depletion microscopy. ‘This analysis revealed major adaptations of intracellular structures that are required to sustain the viral replication and will lead to better understanding of viral pathophysiology and ultimately better medical treatments,’ said Laketa.
These safety ratings are important because such work requires researchers to handle micro-organisms that are potentially hazardous to human health. The IDIP research team conducts its experiments under enhanced biosafety conditions (BSL2 and BSL3) but this adds another layer of complexity to such microscopy-based experiments.
‘There are logistical and technological challenges in setting up microscopy infrastructure in enhanced biosafety conditions to allow maximum flexibility in infectious disease research but at the same time maintain the environment that is maximally safe for everyone involved,’ Laketa explained.
The IDIP research team has also witnessed the ‘dramatically accelerated pace of technical development’ in the world of microscopy. Laketa added: ‘This not only includes hardware developments but the development of other integral components of a microscopy-based experimental approach such as data acquisition and analysis procedures based on machine learning, new fluorophores and close-to-physiological model systems. In IDIP, we continuously need to evaluate new technologies for their applicability in infectious disease research.’
STED developments
Stimulated emission depletion (STED) microscopy is one such emerging super-resolution imaging technique, which has made great strides forward in the field of virology over the last few years. STED is typically the fastest technique for imaging structures below 100nm, making it an ideal candidate for research into small-scale viruses.
In 2014, Professor Stefan Hell won the Nobel Prize in Chemistry for pioneering this technique, where a doughnut-shaped laser beam is used to temporarily silence specific fluorescent molecules. This leads to a smaller spot from where fluorescence is happening and thus super-resolution. A high-resolution picture is then built up, one pixel at a time, by scanning the beams.
Challenges still remain with STED, where the intense light can photobleach the fluorophore. However, Abberior Instruments is now addressing these issues, as Laketa explained: ‘STED has seen a continuous implementation of new technologies since we started using the technique back in 2015. In particular, the photodamage and bleaching have been dramatically reduced with adaptive illumination schemes allowing more super-resolution images to be acquired of the same sample.’
This has led to the development of new, experimental approaches where STED could be used to acquire large 3D stacks or monitor dynamic processes in living cells. ‘There has been a major advancement in making the technology robust and easy-to-use, shaping it into a go-to super-resolution method not much different from standard confocal microscopy in terms of sample preparation requirements and therefore easily employable by laboratories with various degrees of expertise in super-resolution microscopy,’ said Laketa.
The democratisation of STED is an exciting opportunity for researchers and policy makers alike, helping us better understand viruses like Sars-Cov-2 and control their spread.
He also believes there is a ‘big need’ to increase the data efficiency of such cutting-edge microscopy techniques through the introduction of machine learning-based automation. ‘This includes both hardware and software automation in terms of sample positioning, focusing, choosing the right acquisition settings, automatic object detection and subsequent image analysis,’ Laketa concluded.
But STED microscopy is not the end of the chain in super-resolution microscopy. With isotropic resolutions in the range of 1-2 nm, minflux (minimal photon fluxes) microscopy holds great potential to close the resolution gap between STED microscopy and electron microscopy.
At the same time, minflux – in contrast to electron microscopy – is capable of observing cells in their live, native state. The minflux technique will open up completely new fields of research, especially in the field of virus research, but also in other fields, such as mitochondrial research, neuroscience or cancer research.
You can find out more about the advantages of STED and minflux microscopy and its latest contributions to the field of virology in the STED and Virology whitepaper from Abberior Instruments. Information about adaptive illumination is also available in the company’s Adaptive Illumination whitepaper.


