Medical and molecular imaging systems have been around for a long time, and during the 20th century imaging became more well-known for what it could help to reveal in medical applications.
In more recent years the tools to capture and analyse images in the medical and life sciences fields have evolved. Researchers from the universities of Glasgow and Edinburgh recently developed a microscope imaging technique able to record a time-lapse 3D video of a zebrafish heart growing over a day. It is hoped that the work, published in Nature Communications, will provide some insight into how human hearts form, grow and heal.
The fluorescence time-lapse imaging technique uses a thin sheet of laser light to build up images into a full 3D image of the heart layer by layer. The research team combined this with real-time image analysis using visible light to control when to fire the lasers for imaging the heart. Their microscope uses visible light to image the beating heart of the zebrafish through its transparent skin.
By analysing these images, laser light can then be fired selectively at the heart to capture specific moments in its cycle. In this way, exposure to damaging laser light is minimised, allowing the scientists to image the heart over the course of a day without harming the fish. While 3D images of the beating heart had been recorded in the past, the team believes that a time-lapse video of the heart growing had not been possible before this development.
The work was funded by the Royal Academy of Engineering, the Alexander von Humboldt Stiftung, the Engineering and Physical Sciences Research Council, and Amazon.
More recently, researchers at the University of Glasgow have been working on a method of imaging that harnesses AI to turn time into visions of 3D space. Published in Optica, the team were able to make animated 3D images by capturing temporal information about photons, instead of their spatial co-ordinates.
The researchers trained a neural network by showing it thousands of different conventional images of the team moving and carrying objects around the lab, alongside temporal data captured by the single-point detector. Once the network had learned enough about how the temporal data corresponded with the images, it was able to create accurate images from the data alone. In the proof-of-principle experiments, the researchers constructed moving images at about 10 frames per second from the data, although the team believes that the hardware and algorithm used has the potential to produce thousands of images per second.
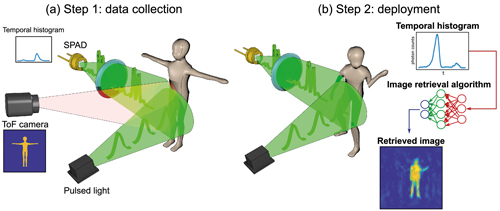
3D imaging with single-point time-resolving sensors. The approach has two steps: data collection and deployment (University of Glasgow; Optica, OSA)
The project was led by Alex Turpin, Lord Kelvin Adam Smith Fellow in data science at the university’s School of Computing Science. The team worked alongside Daniele Faccio and researchers at the Polytechnic University of Milan and Delft University of Technology in The Netherlands.
Turpin noted that one potential application of the technology is to monitor the rise and fall of a patient’s chest in hospital to alert staff to changes in their breathing, or to keep track of their movements to ensure their safety in a data-compliant way.
Turpin and the team are confident that the method can be adapted to any system capable of probing a scene with short pulses and precisely measuring the return echo. ‘This is really just the start of a whole new way of visualising the world using time, instead of light,’ he said.
Currently, the neural net’s ability to create images is limited to what it has been trained to pick out from the temporal data of scenes created by the researchers. But the team believes that, with further training and using more advanced algorithms, it could learn to visualise a much more diverse range of scenes, widening its potential applications in real-world situations.
Turpin said: ‘The single-point detectors which collect the temporal data are small, light and inexpensive, which means they could easily be added to existing systems, like cameras in autonomous vehicles, to increase the accuracy and speed of their pathfinding.’
The next step for the team is to further this research with the development of a portable version of the system with the hope of transferring it to market. But for this, they need some assistance. ‘We’re very excited about the potential of the system we’ve developed, and we’re looking forward to continuing to explore its potential. We’re keen to start examining our options for furthering our research, with input from commercial partners,’ Turpin said.
Share alike
Also bringing their developments into the commercial arena is a team of scientists and students from the University of Sheffield. The researchers have been working on a specialist single-molecule microscope that they believe can be built for a tenth of the cost of current commercially available equipment. Having designed and built the microscope, the team recently shared the blueprint build instructions to help make this equipment available to many labs across the world.
The instructions were published as part of a paper in Nature Communications. The microscope, named the smfBox, is capable of single-molecule measurements. This allows one molecule at a time to be viewed, rather than generating an average result.
This single-molecule method is currently only available at a few specialist labs, because of the cost of commercially available microscopes. The researchers, building on the original smfBox concept, hope to expand the use of single-molecule imaging by giving the wider scientific community detailed build instructions and open-source software to operate the easy-to-use microscope, without the need to invest in expensive infrastructure.
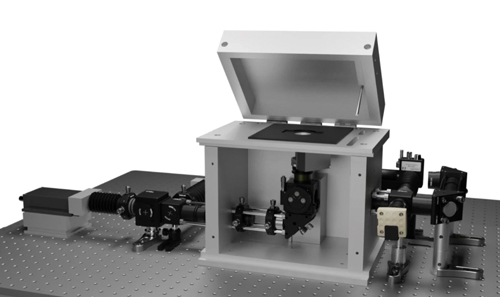

The smfBox is capable of single-molecule measurements. Image: University of Sheffield; Nature Communications
Tim Craggs, the academic lead on the project from the University of Sheffield, explained: ‘We wanted to democratise single-use molecule measurements to make this method available for many labs, not just a few labs throughout the world. This work takes what was a very expensive, specialist piece of kit, and gives every lab the blueprint and software to build it for themselves, at a fraction of the cost.’
The interdisciplinary team spanning the University of Sheffield’s departments of chemistry and physics, and the Science and Technology Facilities Council’s Central Laser Facility, spent a relatively modest £40,000 to build a piece of kit that would normally cost around £400,000 to buy.
The microscope was built with simplicity in mind, so that researchers could use it with little training, while lasers have been shielded in such a way that it can be used in normal lighting conditions. It is made from widely available optics and optomechanical components, replacing an expensive microscope body with machined anodised-aluminium, which forms a light-tight box housing the excitation dichroic, objective, lenses and pinhole.
The university’s Craggs Lab has used the microscope in research to investigate fundamental biological processes, such as DNA damage detection, where improved understanding could lead to better therapies for diseases including cancer. Craggs said: ‘Many medical diagnostics are moving towards increased sensitivity. There is nothing more sensitive than detecting single molecules. In fact, many new Covid tests currently under development work at this level. This instrument is a good starting point for further development towards new medical diagnostics.’
Scanning in 3D
From a commercial perspective, a number of machine vision vendors have been working with laboratories to aid new use cases for the technology. Photoneo’s 3D machine vision technology has been used for scanning the human leg to design and create an ergonomic, high-performance, and also fashionable medical aid – a custom-made orthopaedic orthosis for an ankle joint fracture.
Tomas Kovacovsky, CTO at the company, said: ‘This is unique, as it is generally a challenging task to make 3D scans of the human body in motion. The project started with the presumption that medical aids like orthoses and splints should not only meet anatomical and ergonomic requirements, but they should also correspond to the lifestyle, preferences and taste of the user.’
The aim, therefore, was to create a high-performance, ergonomic and fashionable product. The process combined computational design, 3D scanning and digital fabrication by Subdigital studio, Photoneo, Invent Medical and One3D.
‘The major difficulty,’ continued Kovacovsky, ‘resided in scanning the human body and achieving a precision that would enable the creators to develop a fully ergonomic product.’ This was the first phase of the project, for which Subdigital used Photoneo’s MotionCam-3D.
During the scanning process, the camera moved around the leg to make a 360° scan of the calf and the foot. Photoneo developed a special software for the camera that does not provide a point-cloud but a ‘mesh in real time’ – a 3D model with continuous surface that does not require any post-production.
According to Kovacovsky, this method offers many advantages – especially a high scanning speed and a non-invasive approach. ‘In optimal conditions,’ he said, ‘the 3D model is ready in one or two minutes and the patient does not need to undergo any special procedures before scanning.’
X-factor
Xilinx has been working with institutions such as the (US) National Institutes of Health, Stanford University and MIT to bring a fully functional medical x-ray classification deep-learning model to market, in association with Spline.AI on Amazon Web Services (AWS).
The solution uses an open-source model, which runs on a Python programming platform on a Xilinx Zynq UltraScale+ multiprocessor system-on-chip. This means it can be adapted to suit different, application-specific research requirements. The AI model is trained using Amazon SageMaker and is deployed from cloud to edge using AWS IoT Greengrass. This enables remote machine learning model updates, geographically distributed inference and the ability to scale across remote networks and large geographies.
The open-source design allows for rapid development and deployment of trained models for many clinical and radiological applications in a mobile, portable or point-of-care edge device with the option to scale using the cloud. In fact, the solution has already been used for a pneumonia and Covid-19 detection system, with high levels of accuracy and low inference latency. The development team used more than 30,000 curated and labelled pneumonia images, and 500 Covid-19 images to train the deep learning models. This data is made available for public research by healthcare and research institutes.
Kapil Shankar, vice president of marketing and business development for core markets at Xilinx, elaborated: ‘AI is one of the fastest growing and high-demand application areas of healthcare, so we’re excited to share this adaptable, open-source solution with the industry. As the model can be easily adapted to similar clinical and diagnostic applications, medical equipment makers and healthcare providers are empowered to swiftly develop future clinical and radiological applications using the reference design kit.’
Diverse requirements
Basler has also been developing new technology for use in medical and life science applications. The company has added new models to its Med Ace series, which it introduced at the virtual Analytica trade fair. The USB 3.0 cameras with Sony’s sensitive and high-resolution backside-illuminated sensors IMX178 (6 megapixels) and IMX183 (20 megapixels) come in colour and monochrome versions. Depending on the resolution, the sensors achieve a frame rate of 59 or 17fps, and feature rolling shutter and backside illumination technology.
Peter Behringer, head of product market management for medical at Basler, said: ‘From compact board-level cameras to integrations in point-of-care devices and high-performance 4K60 solutions for surgery, the range is designed to meet all these requirements.
‘Our top priority was to design high-performance and durable cameras that can be easily integrated into medical devices. The cameras also fulfil special requirements, for example, for colour reproduction or cleanliness. We see these requirements in the medical and life sciences area, as well as in factory automation.’
Sponsored: Fluorescence filters for PCR applications
PCR or Polymerase Chain Reaction has been revolutionary in the field of biomolecular sciences, allowing for the study of genesis, morphology, and structure of organisms at the smallest possible scales. PCR is a commonly used DNA amplification method used to replicate low levels of genetic material from various sample types, ultimately replicating the target sequence over and over again. This is a valuable process in a number of applications, for example amplifying small amounts of biological matter from a crime scene.
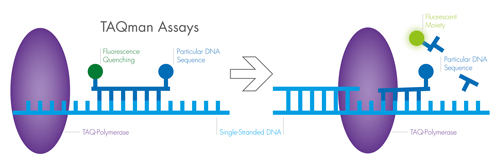
The process begins with the isolation of single stranded DNA (ssDNA); purified double stranded DNA is heated to around 90°C to generate ssDNA by breaking down the hydrogen bonds linking the strands. Next, heat-stable DNA-polymerase (TAQ-Polymerase) is used to replicate the ssDNA chain, beginning the PCR process.
Specific primers help the DNA-polymerase bind and restrict which DNA sequences are replicated. The double-stranded DNA is then heated, and ssDNA is regenerated. The final step involves cooling and primer/probe binding. The ssDNA and primers or probes (described below) bind, and the cycle continues. Each ssDNA is replicated, so the number of copies increases by a factor of two during each round of PCR. The amount of DNA is determined with fluorescent probes.
The fluorescence probes are used to determine reaction efficiency and provide a quantitative analysis of gene expression. This is carried out using either an intercalating dye or hydrolysis probe, which generates fluorescence during PCR under certain conditions. Intercalating dye is introduced to the reaction after the stable polymerase has primed the ssDNA, but before the double strands are replicated. An example dye would be SYBR Green. As the dye binds to the double helix of a DNA strand, it fluoresces more intensely, allowing researchers to correlate brighter fluorescence signals to greater yields of DNA in real time.
Optical filters are a critical component in identifying and monitoring this fluorescence. Laser Components supplies optical filters specifically designed for use in PCR, from partners Omega Optical. Excitation and emission PCR filters are engineered for extremely high transmission over a desired wavelength band, defined by the fluorophore used to probe the reaction. As there is always some spectral overlap between fluorescence probes, it is ideal to use PCR filters with a high optical density out of band, specifically at the region of crossover to achieve the best signal to noise ratio. There are countless different analytical setups for molecular cloning today. From conventional PCR to cutting-edge reverse transcriptase-polymerase chain reactions (RT-PCR), it is now possible to amplify viral RNA as well as DNA. This has proven incredibly useful to virologists studying the sequencing of Sars-CoV-2 – the virus which can cause Covid-19 – in a bid to characterise the virus and to develop novel testing methods.
Our stock PCR filter set is ideal for any fluorescence application using common dyes. For more bespoke application we have extensive custom capabilities.
Alongside our PCR specific filters, we offer a comprehensive range of other fluorescence filter sets. Standard sets are available for over 180 dyes and fluorophores. Long pass emission sets and narrowband sets are available for samples with multiple tags.
Fluorescence sets typically consist of an excitation filter selecting the range of excitation wavelengths, a dichroic mirror and an emission filter blocking the excitation light. The excitation filter is a band-pass which transmits the desired wavelengths and reflects or absorbs the unwanted wavelengths. The transmitted light is reflected through 45° by the dichroic mirror, directing the excitation wavelengths to the sample and allowing the emission to transmit back through. Finally, the emission filter blocks the excitation light and transmits the desired fluorescence to the detector. These filters produced by Omega Optical are characterised by good blocking out of band and high transmission of desired wavelengths to produce a good quality image with a high signal-to-noise ratio.
Filter sets, components, and holders are available for all major microscope manufacturers and models, including Leica, Nikon, Olympus, and Zeiss.


